UTR Sequencing
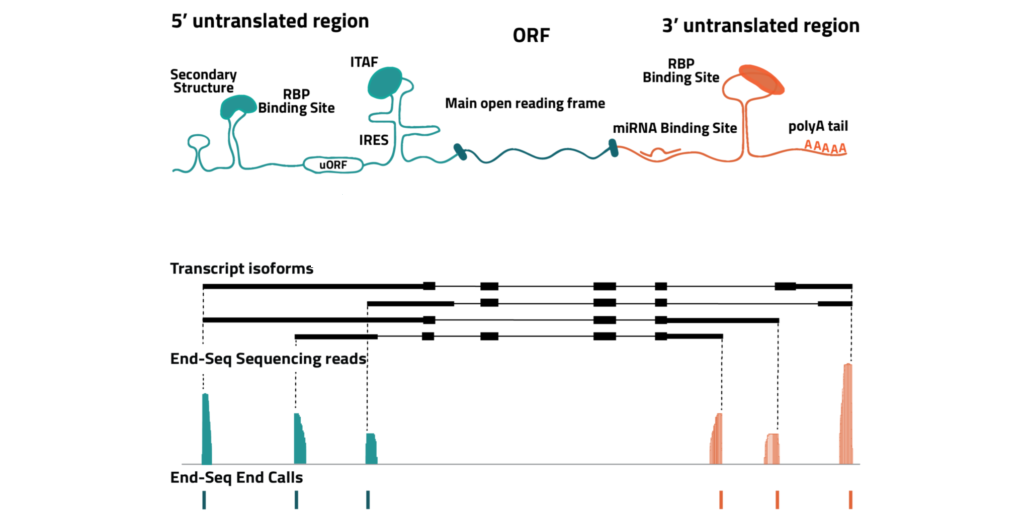
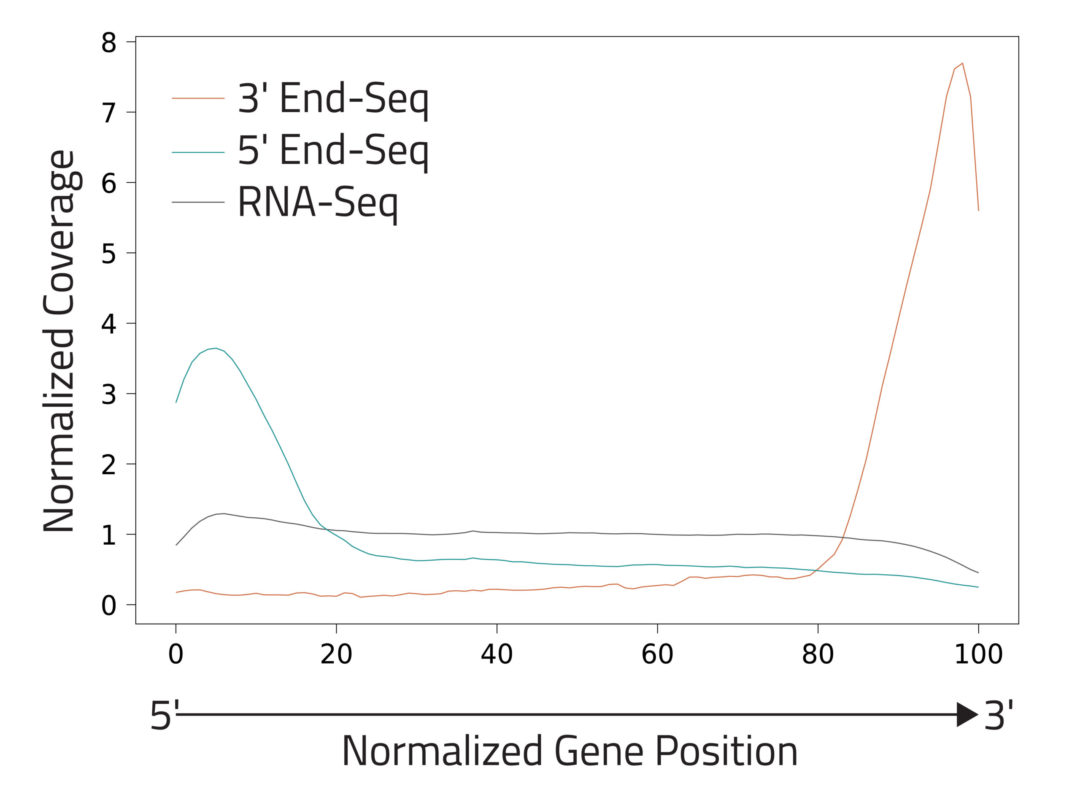
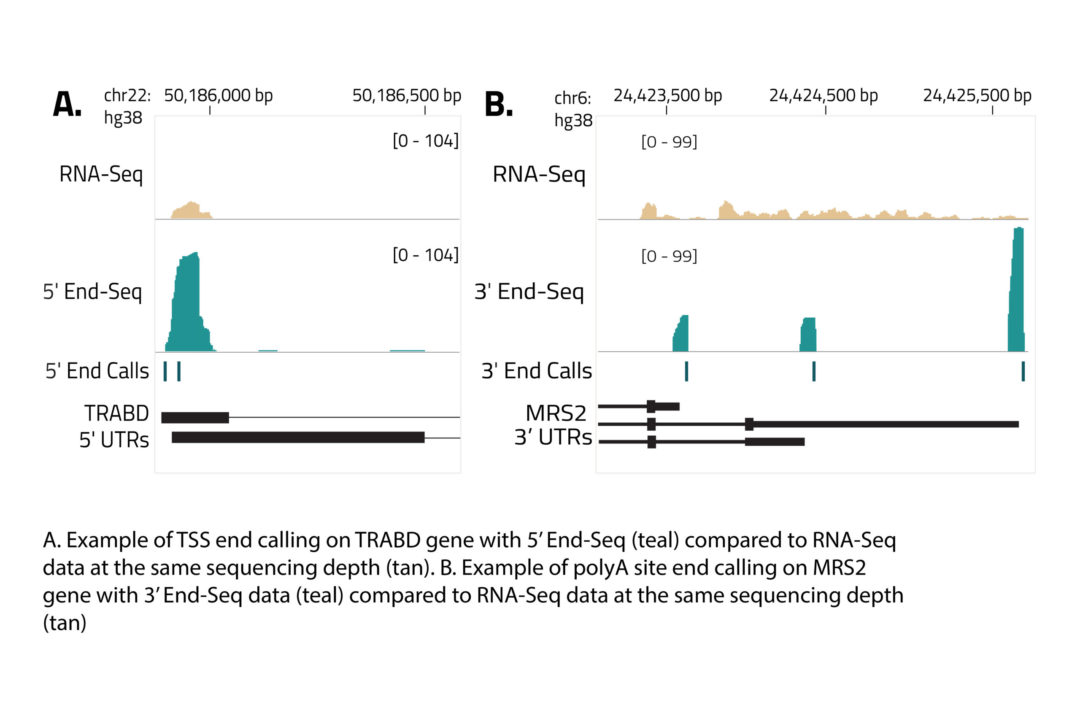
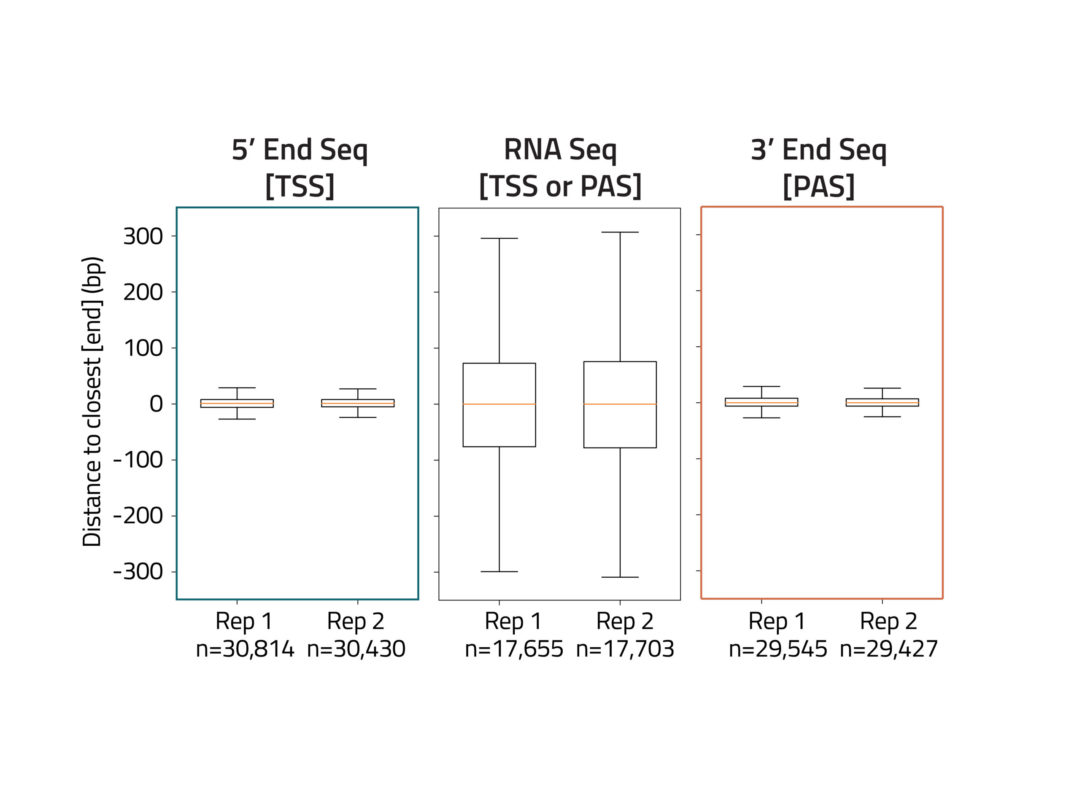
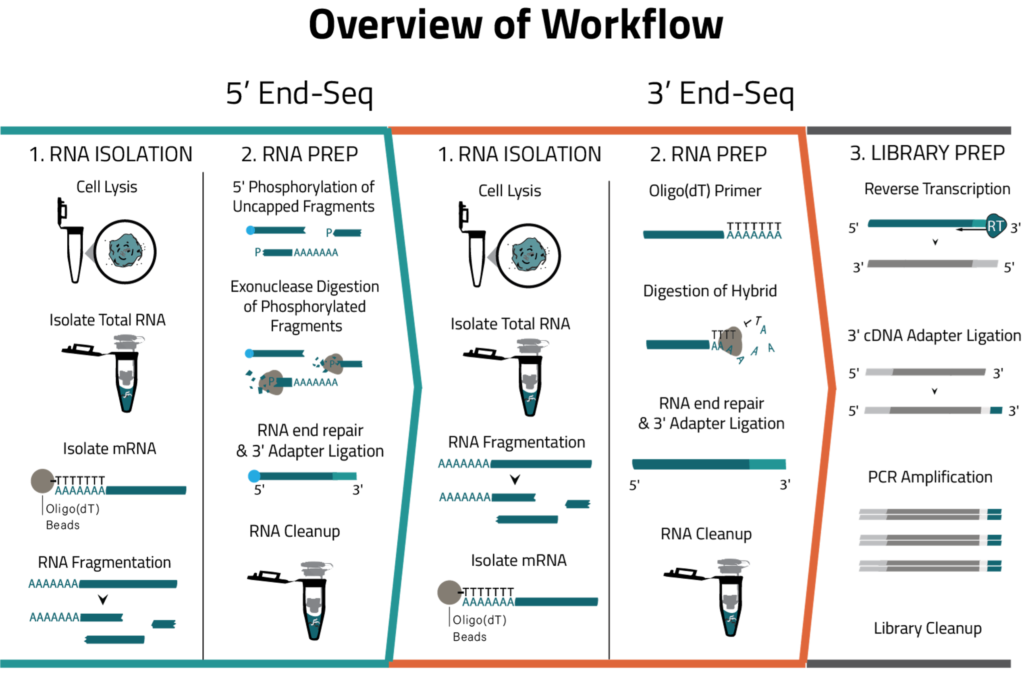
Eclipse BioInnovations´ End-Seq Data Analysis Packages enable the end user to detect all active transcription start sites (TSS) and poly-A-sites, detect novel poly-A-sites with high confidence based on false positive poly-A-site filtering, and further use this data to understand differential gene expression and potentially detect biomarkers in diseased samples. The analysis package uses Fastq files of the raw sequencing reads provided by you. You will receive an HTML report detailing the following data: Single Nucleotide 5´/ 3´End Read Coverage Bedgraph, Single Nucleotide 5´/3´End Calls BED, Filtered Single Nucleotide 5´/3´End Calls (false positive 5´/3´ends are filtered out bioinformatically).
Showing all 4 results