Description
RBP-eCLIP Service: Enhanced CLIP-seq for robust transcriptome-wide identification of RBP targets by NGS
Obtain superior NGS-based target analysis data for your RNA Binding Protein of Interest in cells or tissue provided by you
RBP-eCLIP (Enhanced crosslinking and immunoprecipitation, eCLIP) followed by high-throughput sequencing was developed to provide a robust and reproducible framework to map RNA Binding Protein binding sites on RNAs transcriptome-wide.
Service available for Human (hg38), Mouse (mm10), Rat (rn6), and C. elegans (ce11) genomes.
Service requires an eCLIP-validated antibody (click here for a listing – in case for your RBP no validated antibody is listed, please use the eCLIP Antibody Validation Service )
You provide either 20 Million UV-crosslinked cells or 80 mg flash frozen (not crosslinked) tissue.
Eclipse Bio performs the complete eCLIP-seq protocol in their lab, including enhanced immunoprecipitation, NGS library preparation, sequencing on the Illumina platform (20M reads, SE100), and subsequent comprehensive bioinformatics analysis. RBP eCLIP-seq library preparation comprises an optimized CLIP-seq protocol based on the Van Nostrand, Yeo et al. Nature Methods 2016 paper Robust transcriptome-wide discovery of RNA binding protein binding sites with enhanced CLIP (eCLIP), improving the efficiency of converting immunoprecipitated RNA into high-throughput sequencing libraries
• Highly efficient library pep with low PCR duplication rate: Optimization of enzymatic steps improves library preparation efficiency 1000-fold vs. standard CLIP-seq/iCLIP-seq
• Transcriptome-wide target discovery: RBP binding sites are identified across all regions – exons, introns, UTRs, non coding RNAs, lincRNAs, miRNAs, and retrotransposons
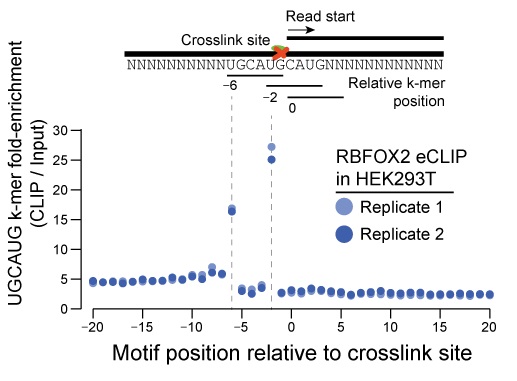
• Precise RBP binding motifs: True in vivo RBP binding sites are identified with single nucleotide resolution
Deliverables:
As final result of your service you receive a comprehensive data analysis package as follows:
RAW SEQUENCING DATA | FASTQ
Reads directly off sequencer and demultiplexed per sample for data back-up and publication
GENOME ALIGNMENTS | BAM
Reads filtered of repetitive elements, aligned, and PCR deduplicated for additional downstream analysis
COVERAGE TRACKS | BIGWIG
Normalized read coverage on positive and negative strands for visualization in a genome viewer such as IGV
SCORED PEAKS | BED
Genomic regions scored after input normalization as putative binding sites (aka “peaks”) for target exploration. Includes log2 enrichment scores and p-values per peak.
BINDING SITE SUMMARY REPORT | HTML
Report for enriched peaks with interactive tables and plots for publication ready graphics. Includes enriched GO terms, KEGG pathways, motifs, and repetitive element analysis.
You can download an example report here
Please contact us to describe your eCLIP-RBP project and to receive a service quotation