Proximo Hi-C Kit for Microbial Samples
1.350,00 €
Description
Proximity Ligation NGS Library Prep from crude microbial samples, for Illumina Sequencing.
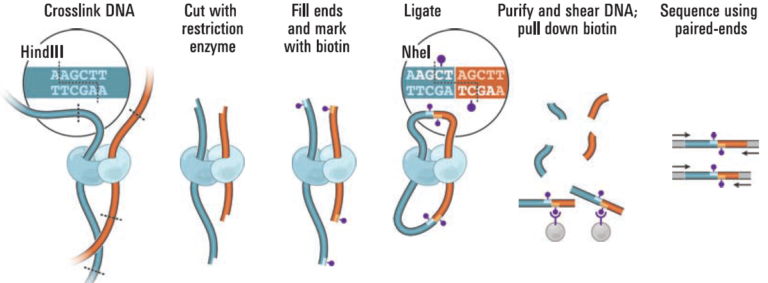
Proximity ligation (Hi-C) libraries are generated from a crude microbial sample. Interactions are captured by crosslinking, digesting, and creating chimeric junctions that are sequenced. The Hi-C sequence data may be used in conjunction with shotgun/whole genome sequence data obtained by standard NGS approaches for metagenome assembly and scaffolding, for TAD calling, as well as for attributing mobile genetic elements such as transposons, phages, plasmids, and antibiotic resistance genes to the host.
• Assemble high-quality metagenomes directly from microbial samples
• No high-molecular-weight DNA or culturing required
• Associate mobile genetic elements (MGEs) such as transposons, phages, plasmids, and antimicrobial resistance genes (ARGs) with their hosts
• User-friendly complete 2-pack Hi-C kit, including reagents for NGS library preparation
• Short-read compatible; yields libraries for Illumina® sequencing
• Individualized data analysis packages for metagenome scaffolding/assembly, and metagenome deconvolution available – please contact us to discuss your project details
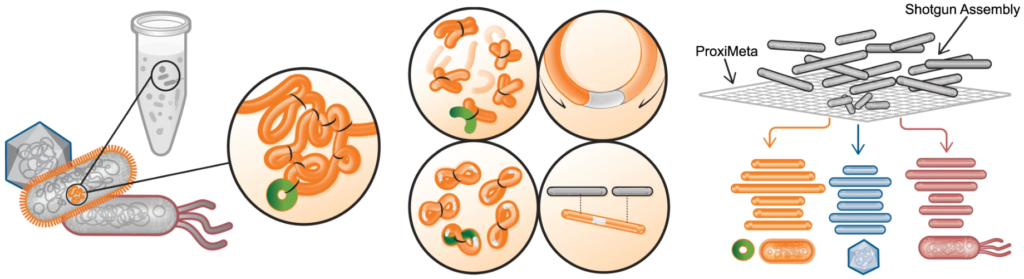
Proximity ligation (Hi-C) libraries are generated from a microbial sample. Interactions are captured by crosslinking, digesting, and creating chimeric junctions that are sequenced and analyzed with a shotgun assembly to deconvolve chromosomes and plasmids into complete genomes.Phase Genomics´ Microbiome Hi-C technology means a true paradigm shift as it enables you to obtain lineage-resolved, complete metagenome-assembled genomes from complex microbial communities.
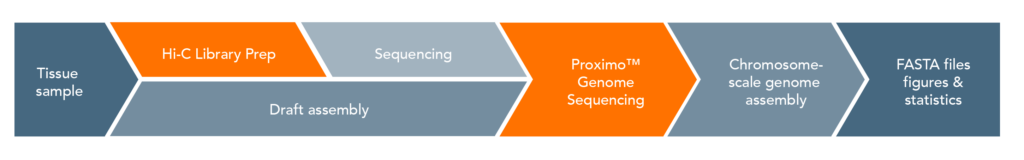
Proximo Workflow:
The proximity ligation (Hi-C) library prep reagents provide a streamlined protocol that does not require culturing of microbes or extraction of high-molecular weight (HMW) DNA. Processing guidelines for different types of metagenomic samples ensure robust results. The 8-step library prep workflow can be completed in 2 working days, and requires only 3 hours of hands-on time. The protocol contains several convenient, safe stopping points.Sequencing may be performed on any Illumina® sequencer. The Proximo Kit yields a dual-indexed proximity ligation library, for which ~50 to 100 million paired-end reads are required. For analyzing your Hi-C sequence data, you also need shotgun/whole genome sequencing data of your samples, which may be performed with a suitable library preparation kit of your choice – contact us for recommendations or to discuss a service project if you do not have prior experience with shotgun/whole genome library preparation and sequencing.
Data Analysis:
Metagenome deconvolution analysis of your sequence data including linkage to plasmids, antibiotics resistance genes, and assignment of phages to their hosts is offered on the ProxiMeta online platform. You can view example reports here. We also offer the ProxiMeta kit and analysis platform, which includes Hi-C reagents for 8 microbiome samples and their online metagenome deconvolution analyses.
Please contact us to let us know your project details and receive a quote.