Description
Enzymatic Fragmentation of Genomic DNA for NGS Applications
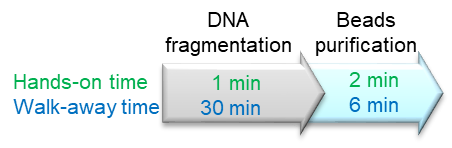
• Fast enzymatic fragmentation: 30-45 minutes, fragment size adjustable by incubation time
• Simple Procedure: 3 minutes of hands-on time, easy handling of many samples at the same time
• Works with both EDTA-free DNA and DNA resuspended in TE buffer
• Ideal for NGS application: random fragmentation, no sequencing bias / consistent sequence coverage
• No capital equipment necessary, no loss of samples
BioDynami´s DNA Fragmentation Enzyme Mix was developed for random fragmentation of genomic DNA (EDTA-free DNA or DNA resuspended in TE buffer) in a fast and simple way. The DNA fragments contain 5´-phosphates and 3´-hydroxyl groups.
Enzyme-based fragmentation of DNA is very simple and effective in comparison to mechanical DNA shearing methods. There are several advantages of enzymatic DNA fragmentation, including easy handling of many samples at the same time, and less loss of samples. Moreover there is no expense for capital equipment necessary.
The resulting DNA fragment size is inversely correlated with the incubation time of step 1 at 20°C. The fragmented DNA can be used for applications such as Next-Generation Sequencing (NGS). BioDynami´s Fragmentation Enzyme does not generate detectable sequencing bias, and sequence coverage is also consistent between enzymatic and mechanical fragmentation.