RNA-Seq Barcode Pooled Libraries
![]()
RNA-Seq Barcode Lentiviral Pooled Libraries link genetic barcode labeling of cells with changes in expression profiles. The approach provides a convenient way to identify cell variations with unique characteristics or biology, and to understand how these groups of variant cells evolve in response to drug treatment, tumor metastasis, or other conditions.
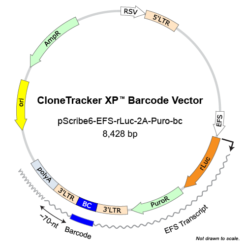
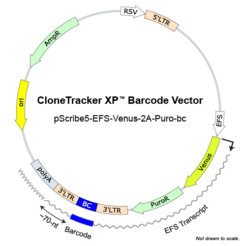
Nucleus Biotech offers CloneTracker XP Barcode Libraries, where the barcode is positioned in the 3’-UTR (or 5′-UTR) of the selection marker so it is expressed as part of the RNA transcript in the cells. As a result, it can be detected by RNA sequencing, as well as DNA sequencing.
You can choose from a 50M XP library, a 10M XP library, and from one to ten XP library pools each of 5M in size.
The 50M XP library is available with either RFP/Puro markers, or RFP/ Thy 1.1 (for mouse in vivo studies)
The 10M XP library can be delivered in 3 different vectors with either red-shifted luciferase (rLuc) derived from Photinus pyralis (Ppy RE9 mutant) as a reporter for bioluminescent imaging (BLI) or Venus Fluorescent Protein/Puro markers.
The ten 5M XP libraries are all in the RFP/Puro vector. To each of the ten pools library specific 14nt barcodes are assigned to enable the differentiation of the 5M pools to each other.
Showing 1–20 of 31 results