Description
The amplicon-based SARS-CoV-2 FLEX NGS panel for Illumina® or Ion Torrent™ is designed for Coronavirus detection, tracking, research and surveillance, enabling complete genome sequencing of the new SARS-CoV-2 virus responsible for the COVID-19 pandemic via amplicon-based target enrichment for Next Generation Sequencing (NGS) on Illumina sequencing platforms. The FLEX panel is built upon the original CleanPlex SARS-CoV-2 Panel and contains strategically designed degenerate primers for robust and confident variant calling even when the virus mutates over time.
• CleanPlex Technology – Ultra high uniformity multiplex PCR and elimination of non-specific PCR poducts allow the ease and flexibility to sequence low input RNA samples for confident infectious disease surveillance and epidemiological research
• High Coverage of Target Regions – 99% coverage of the entire SARS-CoV-2 genome with 343 amplicons in 2 pools, optimized amplification of specific regions in the virus genome for uniform coverage even at low input amount
• High Sensitivity – Can detect down to 1 copy with high confidence compared to 3-5 copies by RT-qPCR
• Fast, Streamlined Workflow – Generate sequencer-ready libraries for Illumina platforms in just 5.5 hours using a simple, four-step protocol with minimal hands on time
• Significant Cost Savings:Only 50K reads per sample required, as high-quality NGS libraries with excellent on-target performance are prepared using CleanPlex technology to enable efficient use of sequencing reads. CleanPlex panels’ on-target rates are usually much higher than those of hybrid captured-based small target enrichment panels
• In Process Control– Human RNA primers are included as library preparation control for confident negative calls
• Degenerate Primers – Strategically designed for consistent coverage to withstand mutations over time
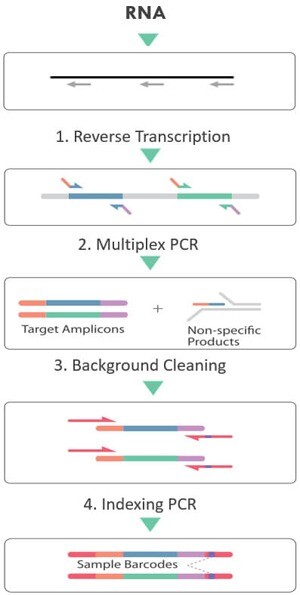
Paragon Genomics´ amplicon-based SARS-CoV-2 FLEX NGS panel is built upon the original CleanPlex SARS-CoV-2 Panel designed as a highly multiplexed target enrichment panel covering the entire genome of SARS-CoV-2 virus (except for 92 bases at the ends). With Paragon Genomics´ CleanPlex technology, the genome of the virus can be amplified from RNA to sequence-ready libraries in 5.5 hours. The panel enables complete genome sequencing, allowing surveillance and epidemiological studies of the new SARS-CoV-2 virus responsible for the COVID-19 pandemic.
The SARS-COV-2 FLEX panel differs from the standard SARS-COV-2 Panel in that it contains degenerate primer designs of polymorphic regions of the genome to allow more robust amplification of variable strains. The FLEX panel also contains a human positive control that amplifies at 100bp longer than the viral library peak to serve as an internal library preparation control, especially for negative samples.
Kit Contents:Reagents for Reverse Transcription and PCR, Multiplex PCR Primers incl. degenerate primers and Human RNA control primers, & CleanPlex Targeted NGS Library Preparation Kit with CleanPlex purification reagents
Additional Products which can be ordered separately:
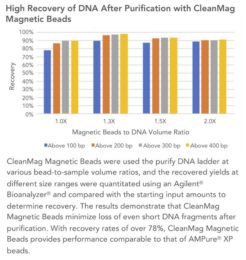
CleanMag® Magnetic Beads – for library purification and size selection
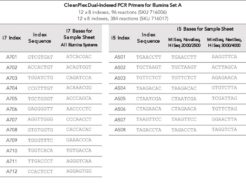
CleanPlex® Dual-Indexed PCR Primers – for multiplexing up to 2688 samples per sequencing run
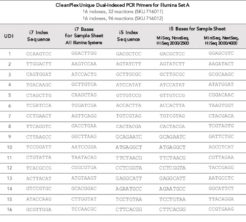
CleanPlex® Unique Dual-Indexed PCR Primers – for low variant calling & other in-depth sequencing applications, and for ultra high multiplexing: each flow cell lane can multiplex up to 384 samples at once; with the multiple flow cell capability on a NovaSeq™ 6000, one can sequence up to 3,072 samples simultaneously for faster turn around and higher throughput workflows.