Description
Universal Spike-In Control RNA Pools for commercial and home-brewed AIRR-seq TCR Profiling Assays based on multiplex RT-PCR or 5’-RACE (SMART) PCR techniques
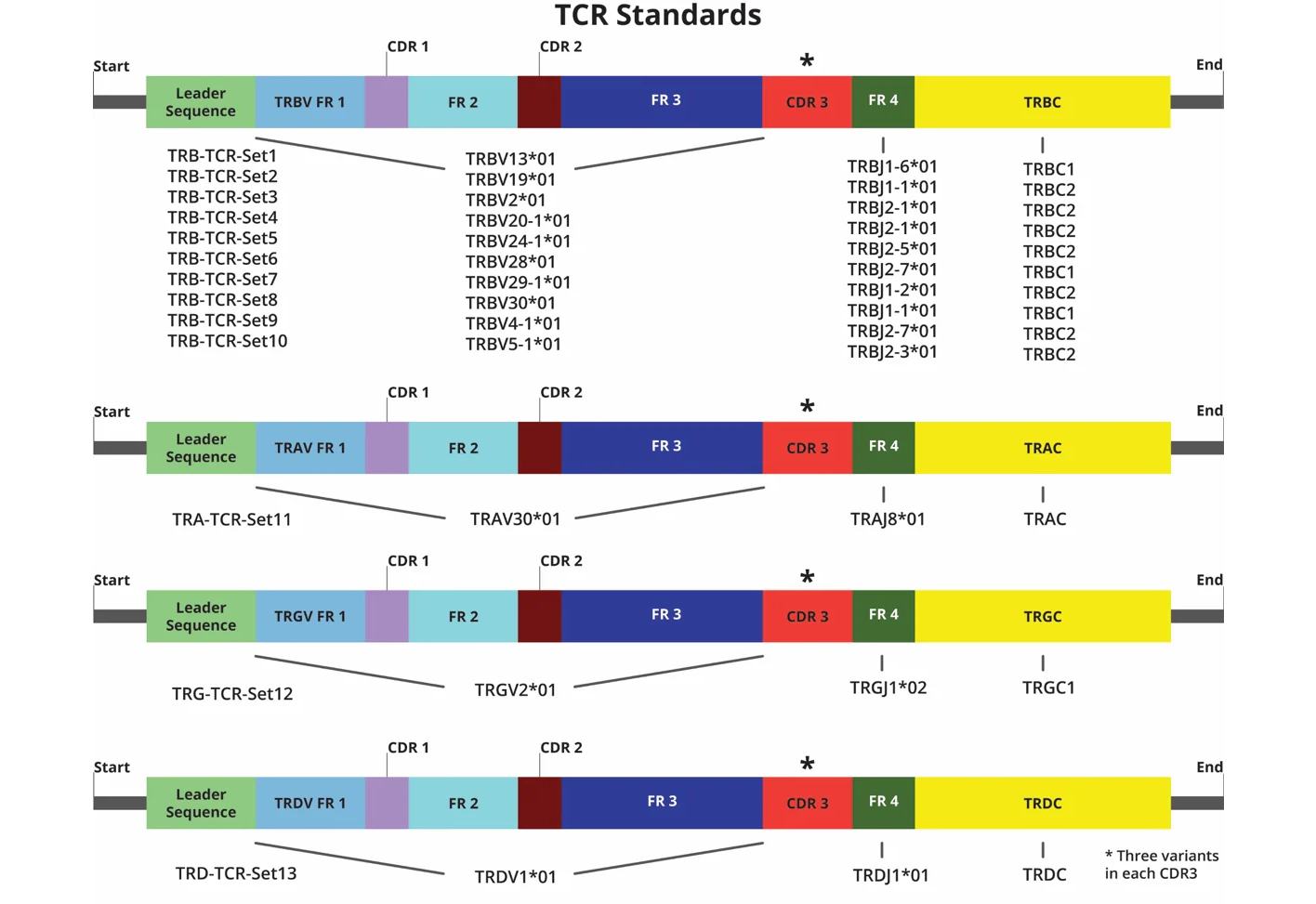
Uniquely available synthetic RNA spike-in control pools for AIRR-seq, designed to ensure consistent and reproducible T-Cell Receptor (TCR) clonotype profiling – contain nearly full-length V(D)JC structures representing all 4 TCR (TRA, TRB, TRD, TRG) chains
The Spike-In RNA Control Pool Set contains 13 different isomer triplex pools of synthetic TCR constructs, each including three constructs with the same V(D)JC isoform receptor structure as featured in the International ImMunoGeneTics information system (IMGT) database, except the CDR3 region which differs from each other by 3 unique single-nucleotide mutations. The three variants in each pool are mixed at a 1:1:1 ratio, each with 2,500 RNA molecules/µl.
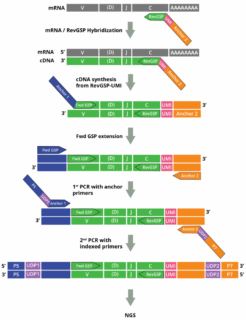
The set of spike-in control RNA pools has been designed to serve as universal RNA positive controls for validating commercial and home brew AIRR Sequencing assays using multiplex RT-PCR such as Cellecta´s DriverMap AIR Kits or 5’-RACE (SMART) assays:
Check Sample Quality: The Spike-In RNA Control Pool Sets may be used to assess sample quality – such as degradation (e.g. in FFPE samples), loss or unbalanced composition of RNA templates, or presence of inhibiting impurities.
Detect Cross Contaminations: Spiking in isoform triplex pools can measure cross-contamination between different samples due to sample mixing, PCR contaminations, or index hopping/jumping in the NGS step.
Product Contents:
13 different TCR isomer pools (50 ul each) of 3 contructs each (1:1:1 ratio) (Typically, 2 µl of isoform control pool should be added to experimental RNA sample)
500ul dilution buffer
5ul PBMC control RNA